Biology Worthy of Life
An experiment in revivifying biology
The Unbearable Wholeness of Beings
This essay is part of a larger work in progress entitled: Toward a Biology Worthy of Life. Original publication: December 9, 2010. Date of last revision: July 4, 2012. Copyright 2010, 2011, 2012 The Nature Institute. All rights reserved.
By clicking on the shaded rectangles at the end of many scientific terms, you can immediately read a definition of the terms in a separate glossary window (or tab, if your browser is set that way).
You can also check out an overview of all the documents associated with this project.
If you try to describe
the living processes of the cell in a rather more living language
than is typically found in the literature of molecular biology — a
language reflecting the artfulness and grace, the well-coordinated
rhythms, and the striking choreography of phenomena such as gene
expression![]() ,
signaling cascades
,
signaling cascades![]() ,
and
mitotic
,
and
mitotic![]() cell division — you will almost certainly hear mutterings about your
flirtation with “spooky, mysterious, nonphysical forces”. You should
expect to hear yourself labeled a “mystic” or — there is no viler
epithet within biology today — a “vitalist”. The previous article in this series reminded one
reader of “some misty Shroud of Turin playing the pan flute and dancing
with the fauns on the meadow”.
cell division — you will almost certainly hear mutterings about your
flirtation with “spooky, mysterious, nonphysical forces”. You should
expect to hear yourself labeled a “mystic” or — there is no viler
epithet within biology today — a “vitalist”. The previous article in this series reminded one
reader of “some misty Shroud of Turin playing the pan flute and dancing
with the fauns on the meadow”.
However grotesque and divorced from serious thought this latter response was, it does distantly reflect a certain sensitivity among biologists — a sensitivity that the nonscientist readily picks up and caricatures. And because this sensitivity is so damaging to the prospects for any profound advance in biological understanding, it deserves to be taken seriously. It was recently given very thoughtful and respectful expression by a first-rank molecular biologist in response to a draft book chapter I had sent him. After describing my views as “very interesting, provocative, and necessary”, and before offering his support for much of what I had to say, he voiced this concern: “You very explicitly dispense with vitalism. Nevertheless, your piece is permeated by an atmosphere that says ‘There is something special about living things’”.
So I believe there is. Animals and plants are a long way from rocks and clouds, and also from automobiles and computers. The need to point this out today is one of the startling aspects of the current scientific landscape. It is true that the concept of “vitalism” has been problematic in the history of biology, but no less so than “mechanism.” The two problems are in fact devilishly intertwined. We will never get straight about vitalism if we do not also get straight about mechanism. And until we sort through the associated confusions, we have little hope of meaningful conversation about many of the perplexities vexing biologists today.
We will see, however, that the shoe is really on the other foot: it’s the conventional literature of biology — and above all the literature of molecular biology — that is steeped in a kind of mysticism now blocking progress. What’s required is a much greater rigor in the use of scientific terminology. And let me add that, in the interest of such rigor, I will avoid as far as possible the use of devil-terms such as “vitalism” and “reductionism” — words that philosophers of biology today generally reject as too ideologically burdened to be of much use. Better to say what one means directly than to lob undiscriminating verbal explosives onto the field of conversation.
Here, then, is my question: Are you and I machines? Are we analyzable without remainder into a collection of mechanisms whose operation can be fully explained, starting from the parts and proceeding to the whole, by the causal operation of physical and chemical laws? It might seem so, judging from the insistent testimony of those whose work is to understand life.
There is little doubt about the biologist’s declared obsession with
mechanisms of every sort — “genetic mechanisms”,
“epigenetic![]() mechanisms”, “regulatory mechanisms”,
“signaling
mechanisms”, “regulatory mechanisms”,
“signaling![]() mechanisms”,
“oncogenic
mechanisms”,
“oncogenic![]() mechanisms”, “immune mechanisms”,
“circadian
mechanisms”, “immune mechanisms”,
“circadian![]() clock mechanisms”, “DNA repair mechanisms”,
“RNA splicing
clock mechanisms”, “DNA repair mechanisms”,
“RNA splicing![]() mechanisms”, and even “molecular mechanisms of plasticity”. The single
phrase “genetic mechanism” yields over 25,000 hits in Google Scholar as I
write, and seems to be rising by hundreds per month. In an informal
analysis of technical papers I’ve collected, I found an average of 7.5
uses of mechanism per article, with the number in a single article
varying from 1 to 32. This is not even counting cognate forms such as
mechanistic and machine.
mechanisms”, and even “molecular mechanisms of plasticity”. The single
phrase “genetic mechanism” yields over 25,000 hits in Google Scholar as I
write, and seems to be rising by hundreds per month. In an informal
analysis of technical papers I’ve collected, I found an average of 7.5
uses of mechanism per article, with the number in a single article
varying from 1 to 32. This is not even counting cognate forms such as
mechanistic and machine.
The odd thing is that I have yet to find a single technical paper in molecular biology whose author thought it necessary to define mechanism or any of the related terms. If the meaning is supposed to be obvious, then presumably we should read the words in a straightforward and concrete way — as indeed seems to be required in the case of molecular machines, which unashamedly projects the human machine shop onto the molecular level. Other usages, however — causal mechanism and mechanistic explanation are examples — evidently convey little more than an idea of physical lawfulness or causation, as when Zaidi et al. (2007) refer to “mechanistic insights into maintenance of cell phenotype through successive cell divisions”. Whatever the implicit definitions may turn out to be, no one will dispute me when I say that the intertwined notions of mechanism and physical law intimately coinhabit the minds of biologists today and are held to be keys for understanding organisms.
But here is the greater curiosity: the same biologists describe the organism in an utterly different manner — so different and yet so seemingly inescapable as to demand, from any thoughtful researcher, some sort of reconciliation with the mechanistic picture. The first thing we will note about this alternative view is its dependence upon a language reaching far beyond that of physics and chemistry.
What Changes at Death?
Anyone whose pet dog has died knows the difference between a living animal and a dead one. Biologists surely know this, too, although (strangely enough!) the difference between life and death does not often figure explicitly in the technical literature presuming to characterize living creatures. You might even think there is something slightly embarrassing about the subject. But, looked at in the right way, the biological literature nevertheless tells us what the biologist knows about the matter. And it is a great deal, even if he would prefer not to admit what he knows.
Think first of a living dog, then of a decomposing corpse. At the moment of death, all the living processes normally studied by the biologist rapidly disintegrate. The corpse remains subject to the same laws of physics and chemistry as the live dog, but now, with the cessation of life, we see those laws strictly in their own terms, without anything the life scientist is distinctively concerned about. The dramatic change in his descriptive language as he moves between the living and the dead tells us just about everything we need to know.
No biologist who had been speaking of the behavior of the living dog will now speak in the same way of the corpse’s “behavior”. Nor will he refer to certain physical changes in the corpse as reflexes, just as he will never mention the corpse’s responses to stimuli, or the functions of its organs, or the processes of development being undergone by the decomposing tissues.
Virtually the same collection of molecules exists in the canine cells during the moments immediately before and after death. But after the fateful transition no one will any longer think of genes as being regulated, nor will anyone refer to normal or proper chromosome functioning. No molecules will be said to guide other molecules to specific targets, and no molecules will be carrying signals, which is just as well because there will be no structures recognizing signals. Code, information, and communication, in their biological sense, will have disappeared from the scientist’s vocabulary.
The corpse will not produce errors in chromosome replication or in any other processes, and neither will it attempt error correction or the repair of damaged parts. More generally, the ideas of injury and healing will be absent. Molecules will not recruit other molecules in order to achieve particular tasks. No structures will inherit features from parent structures in the way that daughter cells inherit traits or tendencies from their parents, and no one will cite the plasticity or context-dependence of the corpse’s adaptation to its environment.
It is a worthwhile exercise: try to think in all these ways about the corpse. You will immediately come up against your experience of the distinction between the dog and its remains, between a strictly physical process and a living performance. Nor need you be ashamed of your experience; the most disciplined biologist, whatever his theoretical inclinations, is leaning very much on the same meanings and distinctions you apprehend. Words such as those cited above, after all, are woven into the decisive explanatory matrix of virtually every contemporary paper in molecular biology — but not in papers dealing with the physical sciences.
Sometimes, in fact, the biologist’s language may reach beyond your own intuitions, as when two researchers say that living organisms not only “issue an integrated response to current conditions” but also “make limited predictions about future environmental changes”, leading to the hope that, with current tools, we can gain “insights into the thought processes of a cell”. The same two researchers describe signaling networks as the “perceptual components of a cell,” responsible for “observing current conditions and making decisions about the appropriate use of resources — ultimately by regulating cellular behaviour” (Hyduke and Palsson 2010). Or you can go back to Barbara McClintock’s Nobel Prize address, when she surmised that “some sensing mechanism must be present . . . to alert the cell to imminent danger”. In the future we should try to “determine the extent of knowledge the cell has of itself, and how it utilizes this knowledge in a ‘thoughtful’ manner when challenged” (McClintock 1983).
But even without references to thought and perception, it’s clear that biologists cannot open their mouths without employing a language of recognition and response, of intention and directed activity, of meaningful information and timely communication, of aberrant actions and corrective reactions, of healthy development leading to self-realization or ill-health leading to death. Yes, all this language sits side by side with the familiar appeals to causal mechanisms. But does it sit comfortably?
We must explore the use of this special language of life — this decidedly non-corpselike language — much further before we can answer that question.
Some Views of the Living Organism
On its face, the language reviewed above — recognize, respond, function, adapt, regulate, and so on — suggests that something is going on over and above a physically lawful performance. In employing the conventional terminology, we describe a kind of directed choreography — a performance whose nature and intent is sufficiently clear for us to judge when errors occur or injury supervenes. (Rocks and clouds do not commit errors or suffer injury.) This implies that we are comfortable making qualitative and aesthetic judgments about health, and can distinguish between coherent and errant meaning in the various informative exchanges continually taking place throughout cell and organism.
We speak, in other words, as though the performer (whatever subject we intend for verbs such as “regulate” and “adapt”) were a real entity or being, capable of signaling or otherwise communicating its own needs and designs, able to make sense of the signals coming from its environment, and, amidst it all, striving to maintain its own distinct, healthy identity.
But it’s not just isolated words and phrases that point to the organism as something more than a collection of physically lawful mechanisms. The larger narratives to which these words lend their meanings are narratives of life, not of carcasses — and much less (as we will see) of machines. Is there any subdiscipline of biology today where research has been reducing cellular processes to a more clearly defined set of causal mechanisms instead of rendering them more ambiguous, more intentional, more plastic and context-dependent, and less mechanical?
We saw in “Getting Over the Code Delusion”
that the
chromosome![]() ,
far from being a kind of fixed, crystalline structure, “is a plastic
polymorphic dynamic elastic resilient flexible
nucleoprotein
,
far from being a kind of fixed, crystalline structure, “is a plastic
polymorphic dynamic elastic resilient flexible
nucleoprotein![]() complex” (Lavelle 2009), and its living — you could almost say
“artistic” — performance is fully as central to its meaning as the
“coded” genetic
sequence
complex” (Lavelle 2009), and its living — you could almost say
“artistic” — performance is fully as central to its meaning as the
“coded” genetic
sequence![]() .
But the chromosome is only one element of the cell. Here are a few of the
many other developing stories in molecular biology that speak of organic
activity fully as dramatic as the temporal and spatial acrobatics of
chromosomes.
.
But the chromosome is only one element of the cell. Here are a few of the
many other developing stories in molecular biology that speak of organic
activity fully as dramatic as the temporal and spatial acrobatics of
chromosomes.
Signaling Pathways.
Signaling pathways![]() are vital means of communication within and between cells. Such pathways
are coherent sequences of molecular interactions by which an initial
encounter — say, the
binding
are vital means of communication within and between cells. Such pathways
are coherent sequences of molecular interactions by which an initial
encounter — say, the
binding![]() of a
hormone
of a
hormone![]() to a cell membrane
receptor
to a cell membrane
receptor![]() — leads to a more or less defined result, or group of results,
“downstream”. One result, for example, might be the activation of a set
of genes.
— leads to a more or less defined result, or group of results,
“downstream”. One result, for example, might be the activation of a set
of genes.
In the conventional machine model of the organism, signaling pathways were straightforward, with a clearcut input at the start of the pathway leading to an equally clearcut output at the end. Not so today, as a team of molecular biologists at the Free University of Brussels found out when they looked at how these pathways interact or “crosstalk” with each other. Tabulating the cross-signalings between just four such pathways yielded what they called a “horror graph”, and quickly it began to look as though “everything does everything to everything” (Dumont et al. 2001). In reality, we see a “collaborative” process that can be “pictured as a table around which decision-makers debate a question and respond collectively to information put to them” (Levy et al. 2010). This directed, corporate decision-making is not the stuff of mere physics and chemistry.
Even considering a single membrane receptor bound by a hormonal or other signal, you can find yourself looking, conservatively, at some two billion possible states, depending on how that receptor is modified by its interactions with other molecules. Despite previous belief, there is no simple binary rule distinguishing activated from deactivated receptors. “The activated receptor looks less like a machine and more like a . . . probability cloud of an almost infinite number of possible states, each of which may differ in its biological activity” (Mayer et al. 2009).
Our problem lies in adequately imagining the reality. When a single protein can combine with several hundred different modifier molecules, leading to practically infinite combinatorial possibilities, and when that protein itself is an infinitesimal point in the vast, turbulent molecular sea of continual exchange that is the cell, and when the cell is one instance of maybe 100 trillion cells of some 250 different major types in the human body, from muscle to bone, from liver to brain, from blood to retina — well, it’s understandable that many researchers prefer not to stare too long at the larger picture. Nevertheless, we should keep in mind that the collaborative process mentioned above involves not just one table with “negotiators” gathered around it, but countless tables with countless participants, and with messages flying back and forth in countless patterns as countless “decisions” are made in a manner somehow subordinated to the unity and multidimensioned interests of the organism as a whole.
In other words, not only are the elements of an individual signaling pathway extremely flexible and adaptive; the individual pathway itself, once thought of as discrete and well-defined, doesn’t really exist — certainly not as a separate “mechanism”. Researchers now speak of the “multi-functionality” or “functional pleiotropism” of signaling nodes, pointing out that signaling networks have “ways of passing physiologically relevant stimulus information through shared channels” (Behar and Hoffmann 2010). More generally, “We tend to talk about pathways and processes as if they are discrete compartments of biology,” write geneticists Emmanouil Dermitzakis and Andrew Clark. “But genes and their products contribute to a network of interactions” — and these interactive networks “differ radically among tissues”.
Whenever we imagine a biological process aimed at achieving some particular result, we need to keep in mind that every element in that process is likely playing a role in an indeterminate number of other significant, and seemingly goal-directed, activities. The mystery in all this does not lie primarily in isolated “mechanisms” of interaction; the question, rather, is why things don’t fall completely apart — as they do, in fact, at the moment of death. What power holds off that moment — precisely for a lifetime, and not a moment longer?
Demise of Lock-and-Key Proteins. Quite apart from its wider context of exchange and interaction, the protein molecule itself is an entire universe of plastic form and possibility. It reminds us that messages do not fly back and forth as disembodied abstractions; they move as dynamically sculptured bodies of force and energy. Their meanings are mimed or gestured — not translated into, or reduced to, a kind of expressionless Morse code.
According to the old story of the machine organism, a
protein-coding![]() DNA
DNA![]() sequence
sequence![]() ,
or
gene
,
or
gene![]() ,
not only specifies an exact messenger
RNA
,
not only specifies an exact messenger
RNA![]() (mRNA) sequence, but the mRNA in turn specifies an exact
amino acid
(mRNA) sequence, but the mRNA in turn specifies an exact
amino acid![]() sequence in the resulting protein, which finally folds into a fixed and
predestined shape. These proteins then carry out their functions by
neatly engaging with each other, snapping into place like perfectly
matched puzzle pieces or inserting into each other like keys in locks.
“There is a sense,” wrote Richard Dawkins, “in which the three-dimensional
coiled shape of a protein is determined by the one-dimensional sequence of
code symbols in the DNA”. Further, “the whole translation, from strictly
sequential DNA ROM [read-only memory] to precisely invariant
three-dimensional protein shape, is a remarkable feat of digital
information technology” (Dawkins 1996, p. 171).
sequence in the resulting protein, which finally folds into a fixed and
predestined shape. These proteins then carry out their functions by
neatly engaging with each other, snapping into place like perfectly
matched puzzle pieces or inserting into each other like keys in locks.
“There is a sense,” wrote Richard Dawkins, “in which the three-dimensional
coiled shape of a protein is determined by the one-dimensional sequence of
code symbols in the DNA”. Further, “the whole translation, from strictly
sequential DNA ROM [read-only memory] to precisely invariant
three-dimensional protein shape, is a remarkable feat of digital
information technology” (Dawkins 1996, p. 171).
We now know how great a misconception this was — a misconception
upon which, in Dawkins’ case, his entire metaphysical-religious-scientific
scheme of the “selfish gene” was erected1. But the truth of the gene and
protein looks quite otherwise than Dawkins’ computerized imagination would
have it. Through
alternative splicing![]() ,
one gene can produce up to thousands of protein variants, while unlimited
additional possibilities arise from
RNA editing
,
one gene can produce up to thousands of protein variants, while unlimited
additional possibilities arise from
RNA editing![]() ,
RNA cleavage
,
RNA cleavage![]() ,
translational regulation, and post-translational modifications.
(“Translation” refers to the process by which an
mRNA
,
translational regulation, and post-translational modifications.
(“Translation” refers to the process by which an
mRNA![]() molecule, along with a large supporting cast, yields a protein.) As for
the finally achieved protein, it need not be anything like the rigid,
inflexible mechanism with a single, well-defined structure imagined by
Dawkins. Proteins are the true shape-changers of the cell, responding and
adapting to an ever-varying context — so much so that the “same”
proteins with the same
amino acid
molecule, along with a large supporting cast, yields a protein.) As for
the finally achieved protein, it need not be anything like the rigid,
inflexible mechanism with a single, well-defined structure imagined by
Dawkins. Proteins are the true shape-changers of the cell, responding and
adapting to an ever-varying context — so much so that the “same”
proteins with the same
amino acid![]() sequences can, in different environments, “be viewed as totally different
molecules” with distinct physical and chemical properties (Rothman 2002,
p. 265).
sequences can, in different environments, “be viewed as totally different
molecules” with distinct physical and chemical properties (Rothman 2002,
p. 265).
Nor is it the case that proteins must choose in a neatly digital fashion between discrete conformations. In contrast to the old “rigid-body” view, researchers now refer to “fluid-like” and “surface-molten” protein structures (Grant et al. 2010; Zhou et al. 1999). Even more radical has been the discovery that many proteins never do fold into a particular shape, but rather remain unstructured or “disordered”. In mammals, about 75% of signaling proteins and half of all proteins are thought to contain long, disordered regions, while about 25% of all proteins are predicted to be “fully disordered” (Uversky 2010). Many of these intrinsically unstructured proteins are involved in regulatory processes, and are often at the center of large protein interaction networks (Gsponer and Babu 2009).
Fluid, “living” molecules do not lend themselves to the analogy with mechanisms, which may explain why the mistaken idea of precisely articulated, folded parts was so persistent, and why the recognition of unstructured proteins has been so late coming. Indeed, this recognition has hardly yet dawned on the biological community as a whole, leading to this lament at a conference on “bioinformatics and bioengineering” at Harvard Medical School:
Experimentalists have been providing evidence over many decades that some proteins lack fixed structure or are disordered (or unfolded) under physiological conditions. In addition, experimentalists are also showing that, for many proteins, their functions depend on the unstructured rather than structured state; such results are in marked contrast to the greater than hundred year old views such as the lock and key hypothesis. Despite extensive data on many important examples, including disease-associated proteins, the importance of disorder for protein function has been largely ignored. Indeed, to our knowledge, current biochemistry books don’t present even one acknowledged example of a disorder-dependent function, even though some reports of disorder-dependent functions are more than fifty years old. (Dunker et al. 2008)
A continuing mechanistic bias is evident even in the negative terms, “disordered” and “unstructured”. The loose, shifting structure of a protein need be no more disordered than the graceful, swirling currents of a river or the movements of a ballet dancer. Given what these proteins harmoniously participate in (among other things: the movements of a ballet dancer) it would be strange to assume that their performance is anything less than graceful and artistic.
The Organism Reveals Itself Through Many Complementary Viewpoints. The living, non-mechanical qualities of the organism are evidenced not only in flexible, collaborative signaling and the plastic dynamism of proteins, but also in the organic unity of the whole, whereby every aspect of the organization is qualified by all the other aspects. There is a mutual interpenetration of processes making it impossible to offer simple chains of causal explanation. The result is that in order to understand the whole we have to take up many different and partial viewpoints — something that was hardly necessary so long as the one-dimensional, machine-like DNA code provided the single and undisputed basis for understanding.
There is, for example, the “ribonome” — the entire collection of
RNA![]() molecules along with the diverse proteins that associate with them.
Australian researcher John Mattick argues that RNA is the true
“computational engine of the cell” (Mattick 2009; Mattick et al. 2009).
This “engine” includes numerous large and small RNAs whose functions are
the result, not simply of their transcription from DNA, but of their
elaborate processing and restructuring within
nucleus
molecules along with the diverse proteins that associate with them.
Australian researcher John Mattick argues that RNA is the true
“computational engine of the cell” (Mattick 2009; Mattick et al. 2009).
This “engine” includes numerous large and small RNAs whose functions are
the result, not simply of their transcription from DNA, but of their
elaborate processing and restructuring within
nucleus![]() and
cytoplasm
and
cytoplasm![]() .
RNA in general
.
RNA in general
is known or strongly implicated to be involved in the regulation of gene expression
(both protein-coding
and noncoding
) at all levels in animals, creating extraordinarily complex hierarchies of interacting controls. This includes chromatin
modification and associated epigenetic
memory, transcription
, alternative splicing
, RNA modification, RNA editing
, mRNA
translation
, RNA stability, and cellular signal
transduction
and trafficking pathways. (Mattick 2007)
It is true that RNA seems to have its hand in just about everything. And
yet others think of
signaling pathways![]() as the decisive integrators: “It is becoming increasingly obvious that
cellular signaling pathways control gene expression programs at multiple
levels, from transcription through RNA processing and finally protein
production” (White and Sharrocks 2010). For still others, chromatin in
general and the
nucleosome
as the decisive integrators: “It is becoming increasingly obvious that
cellular signaling pathways control gene expression programs at multiple
levels, from transcription through RNA processing and finally protein
production” (White and Sharrocks 2010). For still others, chromatin in
general and the
nucleosome![]() in particular provide the clearest vantage point. As structured by
nucleosomes, chromatin “[tells] the story of the genome in a more compact
way without skipping the important features. Well defined, predictive
chromatin signatures offer an elegant framework to comprehensively map all
the functional elements in the human
genome”
in particular provide the clearest vantage point. As structured by
nucleosomes, chromatin “[tells] the story of the genome in a more compact
way without skipping the important features. Well defined, predictive
chromatin signatures offer an elegant framework to comprehensively map all
the functional elements in the human
genome”![]() (Hon, Hawkins and Ren 2009).
(Hon, Hawkins and Ren 2009).
There are other possibilities, too, such as the complex regulation of
protein
translation![]() (Mansfield and Keene 2009). Even the elaborately articulated,
information-rich, and too often overlooked membrane architecture of the
cell can be seen as playing a vital role in orchestrating the production
and proper localization of proteins and, in general, organizing and
structuring the activity of the cell:
(Mansfield and Keene 2009). Even the elaborately articulated,
information-rich, and too often overlooked membrane architecture of the
cell can be seen as playing a vital role in orchestrating the production
and proper localization of proteins and, in general, organizing and
structuring the activity of the cell:
The membranous system of the cell, the backbone of cellular compartmentalization, is the necessary presupposition of its own renewal and replication. Cellular organization in general and membrane-mediated compartmentalization in particular are constitutive of the biological “meaning” of any newly synthesized protein (and thus gene), which is either properly targeted within the context of cellular compartmentalization or quickly condemned to rapid destruction (or cellular “mischief”). At the level of the empirical materiality of real cells, genes “show up” as indeterminate resources . . . If cellular organization is ever lost, neither “all the king’s horses and all the king’s men” nor any amount of DNA could put it back together again (Moss 2003).
Perhaps it is the case that, regardless of the angle or the locale from which we look at the organism, deep inspection will yield a view onto the whole, just as any sentence of a profound and unified text, or any scene of a Greek tragedy, when penetrated deeply enough, opens out onto the meaning of the whole. At the same time, no single view yields a complete or fully adequate description of the whole. There is no one “correct” focus for the biologist; we discover instead numerous complementary perspectives.
Given this broad context, surely we can add the “view from DNA” as another crucially important perspective. And, as biologists have been discovering in recent years, a more contextualized stance opens up a thousand ways to deepen our understanding of DNA itself. This DNA is a participant in the life of the organism, not a dictator of it.
The Organism Is Not a Machine
We come back, then, to that universal preoccupation with mechanistic
terminology mentioned at the beginning. Given the contrast between the
ubiquitous appeal to mechanisms in the technical literature on the one
hand, and the actual qualities of organisms revealed by the language of
biological description on the other, the lack of forthrightness by
researchers regarding what they mean by “mechanism” is remarkable. After
all, there’s no obvious similarity between a sewing machine or clock or
any other machine and, say, a twisting, gesturing
chromosome![]() — or, for that matter, a cat stalking a mouse.
— or, for that matter, a cat stalking a mouse.
I should think that all the foregoing is more than enough to justify wholesale dismissal of the language of mechanism, at least in its more literal meaning. Nevertheless, I will offer here a few further remarks about the impropriety of this language.
The typical living cell is 75-80 percent
water. Its primary activities are flows. Even the parts we have
been taught (by photographs and textbook drawings) to take as fixed
structures are in fact caught up in flows. They themselves are in
one degree or another flows. For example, the filamentous cytoskeleton
that helps give the cell a degree of rigidity and maintain its form “is
not a fixed structure whose function can be understood in isolation.
Rather, it is a dynamic and adaptive structure whose component
polymers![]() and regulatory proteins are in constant flux” (Fletcher and Mullins 2010).
and regulatory proteins are in constant flux” (Fletcher and Mullins 2010).
Moreover, the organism’s relatively fixed structures are themselves the result of flow, not the ultimate cause of it. My favorite example of this comes from my Nature Institute colleague, Craig Holdrege:
Before the heart [in the human fetus] has developed walls (septa) separating the four chambers from each other, the blood already flows in two distinct “currents” through the heart. The blood flowing through the right and left sides of the heart do not mix, but stream and loop by each other, just as two currents in a body of water. In the “still water zone” between the two currents, the septum dividing the two chambers forms. Thus the movement of the blood gives the parameters for the inner differentiation of the heart, just as the looping heart redirects the flow of blood. (Holdrege 2002, p. 12)
The body, it’s been said, is a formed stream. And structures, once stably formed, do not necessarily stay that way. Many of the cell’s membranes are continually yielded up to dissolution and replacement, or they are pinched off to form separate little compartments called vesicles, containing special contents to be delivered somewhere else in the cell before they are dissolved. And the cell as a whole — even a relatively “dead” and undividing cell such as a neuron — may experience a complete replacement of its contents a thousand times or more over the course of its life. Many of the body’s structures are more like standing waves than once-for-all constructed objects.
Mitochondria are energy-supplying organelles, with as few as one or as many as several thousand found in a single cell. The individual mitochondrion is “highly mobile, squirming worm-like back and forth across the cell space to places where energy is needed for special work”. But it often dissolves into fragments, which then fuse with other fragments. “In fact, by placing a cell into a slightly acid medium, all its mitochondria can be made to break up into small spherical beads which, upon return of the cell to normal medium, merge again into strings eventually resuming the appearance and internal structure of a normal mitochondrion” (Weiss 1970, p. 283).
Against the backdrop of context-dependent phenomena such as this, it is
hardly possible to contend that we consist, from the bottom up, of
machine-like devices. The idea is simply disconnected from
reality, reflecting a dogma crystallized from a rarefied mesh of
abstractions rather than an engagement with actual organisms. You might
just as well find “machines” in the currents of a river. When two
scientists write that “Clock genes are components of the
circadian![]() clock comparable to the cogwheels of a mechanical watch” (Albrecht and
Ripperger 2008), it ought to be scandalous. Yet such machine language is
universal, is heavily relied on by otherwise rigorous scientists in their
attempts to explain the organism, has no evident, serviceable meaning, and
working biologists rarely if ever make a serious attempt to justify or
even define it.
clock comparable to the cogwheels of a mechanical watch” (Albrecht and
Ripperger 2008), it ought to be scandalous. Yet such machine language is
universal, is heavily relied on by otherwise rigorous scientists in their
attempts to explain the organism, has no evident, serviceable meaning, and
working biologists rarely if ever make a serious attempt to justify or
even define it.
The parts of a clock are put together in a certain way; the parts of an organism grow within an integral unity from the very start. They do not add themselves together to form a whole, but rather progressively differentiate themselves out of the prior wholeness of seed or germ. They are growing even as they begin functioning, and their functioning is a contribution toward their growing. The parts never were and never are completely separate, never are assembled. A specific bit of food taken in from outside never becomes some new, recognizable part, added to the rest; rather, it is metabolically transformed and assimilated by the ruling unity that is already there. The structures performing this work, such as they are, are themselves being formed out of the work. Does any of this sound remotely like a machine?
When, on the other hand, we do build machines, we impose our designs upon them from without, articulating the parts together so that by means of their external relations they can perform the functions or achieve the purposes we intended for them. Those same relations give us our explanation of the machine’s physical performance. If the behavior of one of the parts depends on internal workings, and if we cannot yet analyze those workings in terms of subparts and their external relations, then we regard the part as a temporarily unexplained “black box”.
One reason we cannot explain the organism through the relations between
parts, is that those parts tend not to remain the same parts from moment
to moment. For example, as most molecular biologists now acknowledge,
there is no fixed, easily definable thing we can call a gene.
Whatever we do designate a gene is so thoroughly bound up with cellular
processes as a whole that its identity and function depend on whatever
else is happening. The larger context determines what constitutes a
significant part, and in what sense, at any particular moment. Where,
then, is any sort of definable mechanism? And the
DNA![]() sequence
sequence![]() is just about the most rigidly fixed element the organism has to offer at
the macromolecular level.
is just about the most rigidly fixed element the organism has to offer at
the macromolecular level.
Certainly there are reasonable analogies between, say, our bones and joints on the one hand and mechanisms such as levers and ball joints on the other. Such analogies can be multiplied many times over throughout the human body. But to avoid falsehood it is necessary to add that these are only approximations.
Bones and joints are not in fact mechanisms. Bones, for example, are continually undergoing an exchange of substances with their environment, and even after the main period of our development is past, they are still being shaped and re-shaped by their use or disuse and by the boundless range of other bodily processes with which they are interwoven. Astronauts on long missions in space lose significant bone mass, density, and strength (Keyak 2009); lions raised in zoos have a bone structure differing from that of lions raised in the wild (Holdrege 1998). It’s certainly true that mechanisms such as ball joints, levers, and cog wheels also suffer change — for example, through wear and tear. But, unlike bones, such mechanisms are not continually re-shaped through the seamless integration of their internal processes with those acting from without. Gears and levers are not maintaining themselves and being maintained in anything like the way an internal organ is.
The pervasive use of the machine metaphor, whether carelessly or by design, imports into biology ideas that have no place there. We have every right to ask the biologist who ceaselessly appeals to mechanisms, machines, and mechanistic explanations, “Please tell us what you mean by these terms”. This doesn’t seem unfair.
Trying to Grasp the Whole Organism
The special nature of biological understanding has been debated for as long as there has been a science of biology, with the debate taking form above all in the long-running dispute, on ever-shifting ground, between mechanists and vitalists. “Mechanism” has meant everything from “the physical organism is a machine, pure and simple” to “the organism is strictly material and is governed by nothing other than physical and chemical processes”. Vitalists, on the other hand, have struggled to glimpse the “special something” that distinguishes living creatures from the non-living, whether it be some physical or quasi-physical “vital force” or simply principles of explanation that cannot be captured in the language of physics and chemistry even if they do not violate physical law.
That these are real issues, rooted both in the apparent distinctiveness of organisms compared to inanimate objects and in our direct awareness of our own life, and that the issues require some kind of resolution that has long escaped the discipline of biology, has been recognized throughout much of the past two centuries. Most biologists in recent decades have vested their hope in what seemed a near-certainty to them: their understanding of the organism would some day be reduced without remainder to the conventional terms of physics and chemistry. The grounds for that certainty having now dramatically dissipated as a result of the molecular disciplines that were supposed to confirm the hopes, any resolution of the long-standing debate seems as remote as ever.
The aspects of the organism triggering the whole dispute have commonly been associated with one or more of the following themes:
The peculiar unity of whole and part: the form, existence, and activities of the parts depend upon, and arise from — are in some sense caused by — the whole, which is therefore expressed in one way or another through every part. It’s much like the relation between individual words and their context — which is not surprising, since language is itself an expression of organic life.
Means-end (“purposive” or “final” or “teleological”) relations: biological activities are carried out as if “with a view toward” or “for the sake of” some end. The organism “aims” to develop and sustain itself as a being with its own particular character. As for the scare quotes employed here: it is agreed on all sides that the directed aspect of biological performance should be distinguished from conscious human purpose, even if the latter is viewed as a coming to intentional self-awareness of whatever expresses itself unreflectively in the wisdom of the body.
The mutual (reciprocal) play of cause and effect: effects are not merely effects, but can simultaneously react back upon their causes. Or, as Kant puts it, the parts “should so combine in the unity of a whole that they are reciprocally cause and effect of each other’s form” (Kant 2000, II.1.65). Perhaps archetypically: as the embryo polarizes into anterior and posterior, each pole is not only “opposite” to the other, but necessarily implied in the other. Each pole is properly formed only by virtue of the other’s being formed. Neither is a unilateral cause of the other.
All three of these features are at least suggested by the rather simpler statement that we find in every organism a meaningful coordination of its activities, whereby it becomes a functioning and self-sustaining unity engaged in a flexible and well-shaped response to the infinitely varying stimuli of its environment. By virtue of this coordination, every local or partial activity expresses its share in the distinctive character of the whole. The ability of the organism to pursue its own ends amid an ever-shifting context means that causal relations become fluid and diffuse, losing all fixity. They are continually subordinated to, or lifted into service of, the agency of the organism as a whole.
There are no doubt many challenges to our understanding in all this, many
issues to be clarified, perhaps even a new language to be worked out. But
the starting point for this effort is clear: governance of the context
over its separate elements, so frequently noted in the literature today,
can be observed at every level, whether we speak of the organism, the
cell, or the
chromosome![]() .
The kind of wholeness we need to reflect upon was well illustrated by
A. E. Boycott in his presidential address to the Royal Society of
Medicine’s pathology section some eighty years ago:
.
The kind of wholeness we need to reflect upon was well illustrated by
A. E. Boycott in his presidential address to the Royal Society of
Medicine’s pathology section some eighty years ago:
We generally think of the blood as something which goes round the body and in so doing brings food to the tissues, takes away their excreta and helps to keep them in communication with one another. But we may also think of it, and sometimes more profitably, as a tissue or organ whose chief business it is to be itself and maintain its own individuality. The blood certainly has a specific structure and a chemical composition, organic and inorganic, which is peculiar to itself. And it shows exquisitely that restorative response to injury which is the chief subject-matter of pathology. Within comparatively narrow limits of natural variations, the volume of blood, the concentration of red cells, the reaction and so on are maintained at steady levels. Though almost every substance which goes into or comes out of the body passes at one time or another through the blood, its composition remains almost constant, and it is this individual characteristic which entitles us to have “normal” standards of haemoglobin, red cells and the rest. All experience shows, too, that it is very difficult experimentally to produce deviations from these normal values of more than a fleeting character, and under a great variety of circumstances the blood persists in remaining itself. (Boycott 1929)
“Persists in remaining itself”. The phrase may not quite rest comfortably with modern scientific sensibilities. Nor is it the only such phrase. But reasonable interpretations have long been on offer, as we will now see.
What Does “More Than the Sum of Its
Parts” Mean? Variation of the parts amid
relative constancy of a well-ordered whole that strives to remain itself:
this was a central theme of one of the most prominent and now most
unjustly neglected scientists of the past century. By all accounts a
distinguished cell biologist, Paul Weiss pursued active research from the
1920s on into the 1970s, when he was awarded the National Medal of
Science. He pioneered many techniques of tissue culture while pursuing
important work in neurobiology,
morphogenesis![]() ,
limb and nerve regeneration, and cell
differentiation
,
limb and nerve regeneration, and cell
differentiation![]() .
His awards and recognitions were many.
.
His awards and recognitions were many.
Before coming to America, Weiss received an “old-school” education in Austria, which may account for the fact that he was aware of certain broader issues in biology from the very outset of his career. A scientist’s scientist in terms of his mathematical, experimental, and observational rigor, he couldn’t help noticing organismal behavior that didn’t fit the prevailing mechanistic models. For example, his powerful arguments against the gene-centered understanding of the organism, which we will touch on below, were founded on the most basic facts of observation and the most straightforward, unassailable reasoning — and they were arguments that would today be widely accepted. But at the time his was a beleagured voice; the almost arrogant confidence of molecular biologists, founded on unyielding philosophical commitment to the explanatory hegemony of the gene, prevented them from taking in and digesting his arguments. But now, if I’m not mistaken, there is a reawakening interest in what this rather low-key and incisive prophet had to say.
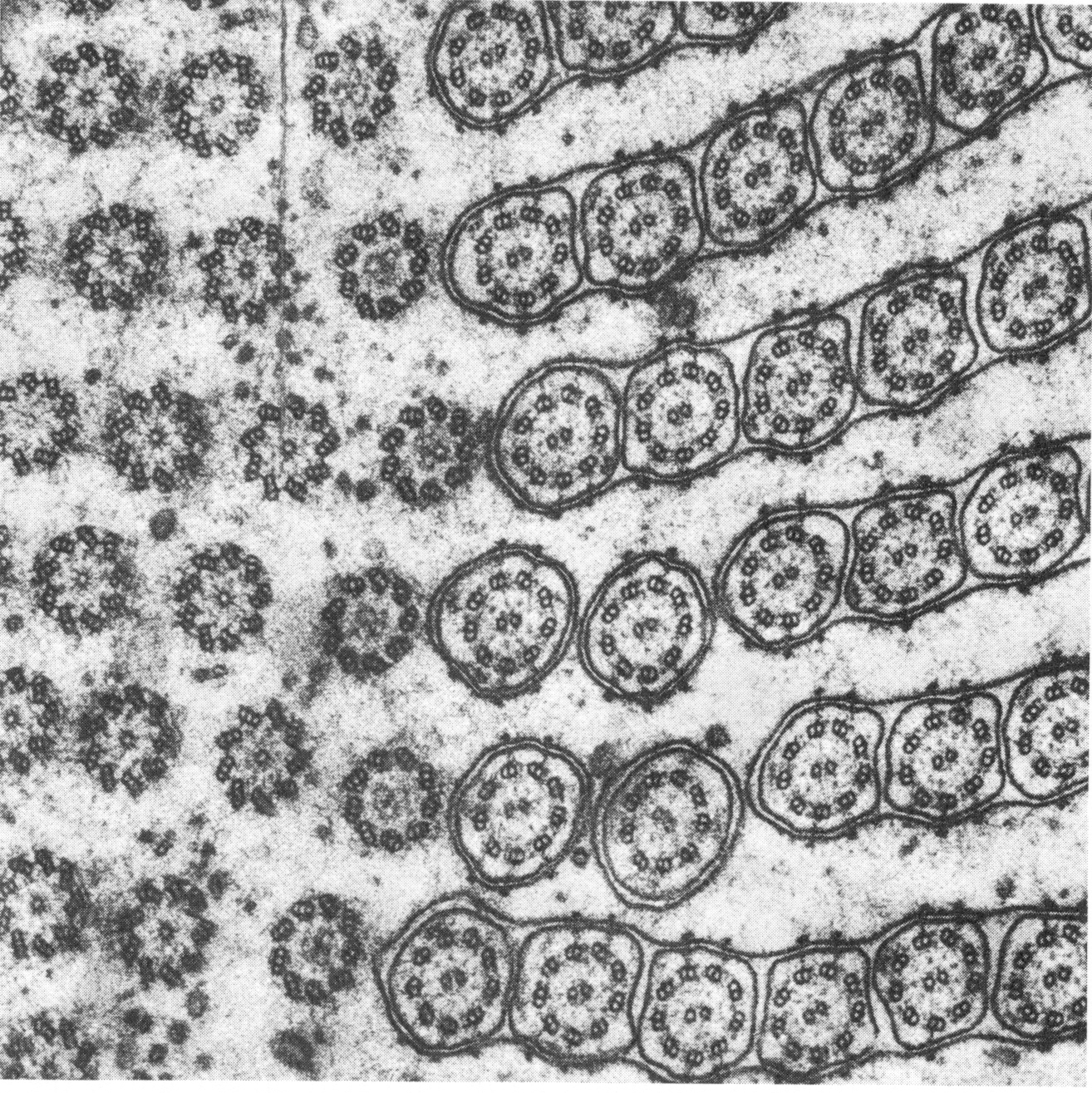
Picking up the theme of Boycott about the constancy of the blood amid change, Weiss provided numerous examples of global unity and harmony superimposed upon lower-level variation (for example, Weiss 1959; 1962; 1967; 1970). One is the electron micrograph below, which shows a tangential section grazing the surface of a single-celled ciliate protozoan. Because the angle of the section is slightly oblique, the circular structures — each one a single cilium with eleven parallel fibers (nine in a circle and two in the middle) — are shown cut at varying depth, revealing different aspects of the structures. The placement and form of all the details shows no constancy. And yet that unevenness, which might be expected to lead to ever less order in the overall composition, is nevertheless disciplined toward a larger, patterned harmony.
| ||
| From Weiss 1962. Attributed to I. Gibbons. |
Weiss shows repeatedly in his various analyses that the mechanical forces or physical dimensions or one-to-one interactions at the level of the parts of an organism are inadequate to determine the coherence of the scheme into which the parts are fitted. We can’t compare the arrangement of cilia shown above to the way rigid, precisely shaped bricks can be laid out in a pattern determined by their shapes. We see “certain definite rules of order” that “apply to the dynamics of the whole system . . . reflected in the orderliness of the overall architectural design, which cannot be explained in terms of any underlying orderliness of the constituents” (Weiss 1970, p. 286).
Much the same applies to the
pluripotent![]() cells of the very young embryo. A given cell can be moved from one place
to another, resulting in a completely different fate for that cell within
the developing organism. This indicates that the cell’s fate is
determined “on the fly”: a governing dynamic disposes of each part
according to the needs of the overall pattern. The developing relations
between the individual cells are more a result of than a cause of the
order of the whole.
cells of the very young embryo. A given cell can be moved from one place
to another, resulting in a completely different fate for that cell within
the developing organism. This indicates that the cell’s fate is
determined “on the fly”: a governing dynamic disposes of each part
according to the needs of the overall pattern. The developing relations
between the individual cells are more a result of than a cause of the
order of the whole.
Evidently, besides its full complement of “genetic information”, each cell needs still additional “topical information” derived from the field structure of the collective mass. How otherwise could any unit know just what scrap from its full grab bag of inside information to put to work at its particular station in order to conform to the total harmonious program design? Clearly, left solely to their own devices, the individual cells and their entrapped genomes
would be as incapable of producing a harmonious pattern of development
as a piano with a full keyboard would be of rendering a tune without a player. (Weiss 1973, p. 35)
It is crucial to realize what Weiss is not saying. He is not saying that the laws of physics are violated in the formation of organic patterns. He himself spent many years elucidating the play of physical forces in such situations. What is being coordinated is nothing other than this play of forces. His point is that, whatever the level we analyze, from macromolecular complexes, to organelles, to cells, to tissues, to individual organs, to the organism as a whole, we find the same principle: we cannot reconstruct the pattern at any level of activity by starting from the parts and interactions at that level. There are always organizing principles that must be seen working from a larger whole into the parts.
Weiss expressed this truth in a significant formula:
VS < Σ(va + vb + vc + . . . vn)
That is, if you take a given fraction, A, of an organic complex such as a cell, and if you measure all the fluctuations of its physical and chemical parameters over a period of time, summing them as va, and if you do the same with all the other fractions, B, C . . . and if, finally, you record all the measurable features of the total complex, S, as a whole, along with their variations, then you reach this conclusion: the total variance of all the subprocesses within a cell (jumping up a level, you could also say: “the total variance of all the cells in a tissue”) is less than the sum of all the variances of the individual subprocesses (or separate cells in the tissue). “The formula represents an ‘operational’ description of what it is that makes the cell as a unity ‘more than the sum of its parts’” (Weiss 1963, pp. 395-6).
In other words, despite the countless processes going on in the cell, and despite the fact that each process might be expected to “go its own way” according to the myriad factors impinging on it from all directions, the actual result is quite different. Rather than becoming progressively disordered in their mutual relations (as indeed happens after death, when the whole dissolves into separate fragments), the processes hold together in a larger unity. The behavior of the whole “is infinitely less variant from moment to moment than are the momentary activities of its parts”:
Small molecules go in and out, macromolecules break down and are replaced, particles lose and gain macromolecular constituents, divide and merge, and all parts move at one time or another, unpredictably, so that it is safe to state that at no time in the history of a given cell, much less in comparable stages of different cells, will precisely the same constellation of parts ever recur . . . Although the individual members of the molecular and particulate population have a large number of degrees of freedom of behavior in random directions, the population as a whole is a system which restrains those degrees of freedom in such a manner that their joint behavior converges upon a nonrandom resultant, keeping the state of the population as a whole relatively invariant. (Weiss 1962, p. 6)
We might say that a given type of cell (or tissue, or organ, or organism) insists upon maintaining its own recognizable identity with “unreasonable” tenacity.
It turns out, then, with a touch of irony, that less change is what shows the whole cell to be more than the sum of its parts. It’s as if there were an active, coordinating agency subsuming all the part-processes and disciplining their separate variabilities so that they remain informed by the greater unity. The coordination, the ordering, the continual overcoming of otherwise disordering impacts from the environment so as to retain for the whole a particular character or organized way of being, expressively unique and different from other creatures — this is the “more” of the organism that cannot be had from the mere summing of discrete parts. The center holds, and this ordering center — this whole that is more than the sum of its parts — cannot itself be just one or some of those parts it is holding together. When the organism dies, the parts are all still there, but the whole is not.
Animistic Impulses in Biology.
Or consider
DNA![]() and the vast array of proteins and other molecules that must cooperate
with it in all its functions. One of the most stupefying blind spots of
the molecular biology era notwithstanding, a DNA molecule by itself is
without meaning for the organism; it cannot do anything. As
Harvard biologist Richard Lewontin once wrote, it is “a dead molecule,
among the most nonreactive, chemically inert molecules in the living
world” (Lewontin 1992). Its meaning is as much a function of the
molecules with which it interacts as it is a property of its own
structure. Or, in Weiss’ words: the irreducible truth is that “Life is a
dynamic process. Logically, the elements of a process can be only
elementary processes, and not elementary particles or any
other static units” (Weiss 1962, p. 3).
and the vast array of proteins and other molecules that must cooperate
with it in all its functions. One of the most stupefying blind spots of
the molecular biology era notwithstanding, a DNA molecule by itself is
without meaning for the organism; it cannot do anything. As
Harvard biologist Richard Lewontin once wrote, it is “a dead molecule,
among the most nonreactive, chemically inert molecules in the living
world” (Lewontin 1992). Its meaning is as much a function of the
molecules with which it interacts as it is a property of its own
structure. Or, in Weiss’ words: the irreducible truth is that “Life is a
dynamic process. Logically, the elements of a process can be only
elementary processes, and not elementary particles or any
other static units” (Weiss 1962, p. 3).
“But”, comes the reply, “all the molecules involved in these processes are made by DNA”.
That’s not true. First, as I just mentioned, DNA by itself cannot make
anything. Second, many crucial molecules that shape the functioning of
the cell, including all lipids and carbohydrates, do not derive from DNA.
This reminds us that the central functioning of metabolism — the
transformation of nutrients in the cell — is not in any realistic
sense controlled by DNA. The reverse is just as true; metabolic
processes send
signals![]() to DNA when its resources are wanted. Third, the proteins and
noncoding
to DNA when its resources are wanted. Third, the proteins and
noncoding![]() RNAs
RNAs![]() that do derive from DNA are extensively and significantly modified by
processes in the
cytoplasm
that do derive from DNA are extensively and significantly modified by
processes in the
cytoplasm![]() ,
with their functions depending heavily on these modifications. Fourth,
the enzymes and other proteins essential for
transcribing
,
with their functions depending heavily on these modifications. Fourth,
the enzymes and other proteins essential for
transcribing![]() DNA certainly cannot be described as mere products of DNA because they are
part of the essential means for producing all proteins, including
themselves. And fifth, DNA, far from being responsible for everything in
the cell, is itself the responsibility of the cell, which goes through a
balletic drama of scarcely conceivable complexity in order to replicate
and preserve this vitally important molecule.
DNA certainly cannot be described as mere products of DNA because they are
part of the essential means for producing all proteins, including
themselves. And fifth, DNA, far from being responsible for everything in
the cell, is itself the responsibility of the cell, which goes through a
balletic drama of scarcely conceivable complexity in order to replicate
and preserve this vitally important molecule.
In sum: all cellular constituents, including DNA, originate from the cell and organism as a whole.
To say, as Nobel laureate Max Delbrück once did, that DNA could be conceived in the manner of Aristotle’s First Cause and Unmoved Mover, since it “acts, creates form and development, and is not changed in the process” (Delbrück 1971) — well, that’s the stupefying blind spot, a blind spot that to one degree or another ruled the entire era of molecular biology through the turn of the current century. It was already recognized and warned against by the German botanist F. Noll in 1903, who pointed out how (in E. S. Russell’s paraphrase) “the chief theorists have tried to solve the problem of development by assuming a material and particulate basis [today’s ‘gene’], without however attempting to explain how the mere presence of material elements could exert a controlling influence on development. They have been forced to ascribe to such abstract material units properties and powers with which they would hesitate to credit the cell as a whole” (Russell 1930, p. 287).
Weiss emphasizes very much the same point: because there is no possible way to make global sense of genes and their myriad companion molecules by remaining at their level, researchers have “simply bestowed upon the gene the faculty of spontaneity, the power of ‘dictating’, ‘informing’, ‘regulating’, ‘controlling’, etc.” (Weiss 1970, p. 302). And today, one could add, there’s at least an equal emphasis on how other molecules “regulate” and “control” the genes! Clearly something isn’t working in this picture of mechanistic control. And the proof lies in the covert, inconsistent, and perhaps largely unconscious invocation of higher coordinating powers through the use of these loaded words — words that owe their meaning ultimately to the mind, with its power to understand information, to contextualize it, to regulate on the basis of it, and to act in service of an overall goal.
Weiss considers terms such as “regulate”, “organize”, and “control” an “obvious reversion in modern guise to animistic biology, which let animated particles under whatever name impart the property of organization to inanimate matter” (Weiss 1962). He says this, not because he resists the ideas of regulation, organization, and so on, but rather because he refuses to ascribe the power of regulating and organizing to specific material parts of the organism, which then gain a kind of magical quality. Whatever regulates a set of interacting parts cannot be found in one of the parts being regulated. To see the principles of regulation governing any set of parts, we have to step back, or up, until we can recognize a unity and harmony that operates, so to speak, between the parts, becoming visible only from a more comprehensive, relational vantage point.
This unity and harmony may represent a genuine difficulty for our understanding, if only because few in recent decades have bothered to address it. But until we see the problem where it actually lies, instead of concealing it in molecules with mystical qualities, we can hardly begin the work of trying to understand.
As I noted earlier, serious researchers have long recognized the “problem” of biological explanation. It’s just that the issues were largely set aside in the era of molecular biology due to the expectation that they were now completely manageable and well on their way to routine solution. Biology would soon be rid of its troublesome language of life in favor of well-behaved molecular mechanisms. And yet today, after several decades of stunning progress in molecular research, it is no more possible than it was two hundred years ago to construct a single paragraph of properly biological description that does not draw on a meaningful language of living agency considered improper in chemistry or physics.
If we want to reckon with the holism, the coordination and organization, the means-end relationships that are continually appealed to in biological explanation, one way forward might be to take the biologist’s persistent language — minus its mystical tendencies — seriously and at face value. Perhaps the biologist describes what she actually sees, and perhaps the living qualities of the organism are not really as spooky as they are sometimes made out to be. Perhaps it never did make sense to try to understand the world from the bottom up, never made sense to dismiss the richest, most multi-faceted phenomenal displays — the most organically unified realizations of the world’s creative potential, such as we find in the performance of whole living creatures — as if they were, by very reason of the fullness of their revelation, the most unreal and misleading guides to the true nature of things.
Who Regulates the Regulators?
Before concluding, it remains only to illustrate ever so briefly what happens when you mix the language of organic coordination with that of mechanistic control. It’s not a pretty sight. A very worthy paper that landed in my email box while I was writing the preceding paragraph serves as well as any to illustrate the situation. It concerns the p53 protein:
The tumor suppressor p53 is a master sensor of stress that controls many biological functions, including [embryo] implantation, cell-fate decisions, metabolism, and aging . . . Like a complex barcode, the ability of p53 to function as a central hub that integrates defined stress signals
into decisive cellular responses, in a time- and cell-type dependent manner, is facilitated by the extraordinary complexity of its regulation. Key components of this barcode are the autoregulation loops, which positively or negatively regulate p53’s activities.
We have, then, a master sensor (p53) that controls various fundamental cellular processes, and yet is itself dependent on the signals it receives and is subject to “extraordinarily complex” regulation by certain autoregulation loops. While all these loops regulate p53 (some positively and some negatively), one of them, designated “p53/mdm2,”
is the master autoregulation loop, and it dictates the fate of an organism by controlling the expression
level and activity of p53. It is therefore not surprising that this autoregulation loop is itself subject to different types of regulation, which can be divided into two subgroups . . . (Lu 2010)
So the master controlling sensor is itself subject to a master controlling process (one of several regulatory loops) that dictates the fate of the organism. But this master loop, it happens, is in turn regulated in various manners (as the author goes on to say in the rest of the article) by a whole series of “multi-layered” processes, including some that are themselves “subject to direct regulation by mdm2” — that is, they are regulated by an element of the regulatory loop they are supposed to be regulating.
I can hardly begin to describe the stunning complexity surrounding and supporting the strikingly diverse performances of the p53 protein. But by now every biologist knows how such “regulatory” processes extend outward without limit, connecting in one way or another with virtually every aspect of the cell. The article on p53 makes an admirable effort to acknowledge and summarize the almost endless intricacy and contextuality of p53 functioning and, with its language of mechanism and control, it does not differ from thousands of other papers. But that only underscores the undisciplined terminological confusion continuing to corrupt molecular biological description today. When regulators are in turn regulated, what do we mean by “regulate” — and where within the web of regulation can we single out a master controller capable of dictating cellular fates? And if we can’t, what are reputable scientists doing when they claim to have identified such a controller, or, rather, various such controllers?
If they really mean something like “influencers,” then that’s fine. But influence is not about mechanism and control; the things at issue just don’t have controlling powers. What we see, rather, is a continual mutual adaptation, interaction, and coordination that occurs from above. What we see, that is — once we start following out all the interactions at a molecular level — is not some mechanism dictating the fate or controlling an activity of the organism, but simply an organism-wide coherence — a living, metamorphosing form of activity — within which the more or less distinct partial activities find their proper place. The misrepresentation of this organic coherence in favor of supposed controlling mechanisms is not an innocent inattention to language; it’s a fundamental misrepresentation of reality at the central point where we are challenged to understand the character of living things.
(Here is another example — drawn at random from countless such cases — showing misuse of the language of control.)
How the organism holds together and makes sense is surely what the employers of such language are really trying to capture. One sympathizes with them. The problem is that their science gives them a respectable (and extremely valuable) language of analysis, while it’s still stumbling around looking for a language able to comprehend unities or wholes — a “systems” language, some would say. The difficulty is owing to the stubborn proviso that this language mustn’t come too uncomfortably close to infringing the taboo against recognizing mind and meaning, direction and intention, lest the world become unsafe for objects and mechanisms. So the researcher is left with an interesting problem: to make sense of the organism without finding any real meaning in it — least of all the meaning traditionally associated with living beings. Systems may perhaps be tolerated; at least they are reassuringly vague and anonymous, and invite casual manipulation. But who knows what disagreeable entanglements might follow once we find ourselves staring into the face of other beings?
Notes
1. On the religious dimension of Dawkins’ work, see the late Australian philosopher David Stove’s somewhat eccentric book, Darwinian Fairytales (1995), especially chapters 7, 9, and 10.
References
Albrecht, Urs and Jürgen A. Ripperger (2008). “Clock Genes”, available at http://unifr.ch/biochem/assets/files/albrecht/publications/AlbrechtRipperger.pdf. [Accessed Dec. 6, 2010.]
Behar, Marcelo and Alexander Hoffmann (2010). “Understanding the Temporal Codes of Intra-cellular Signals”, Current Opinion in Genetics and Development vol. 20 (Dec.), pp. 684-93. doi:10.1016/j.gde.2010.09.007
Boycott, A. E. (1929). “The Blood as a Tissue: Hypertrophy and Atrophy of the Red Corpuscles”, Proceedings of the Royal Society of Medicine vol. 23, no. 1 (Nov.), pp. 15-25.
Dawkins, Richard (1996). The Blind Watchmaker: Why the Evidence of Evolution Reveals a Universe without Design. New York: W. W. Norton.
Delbrück, M. (1971). “Aristotle-totle-totle”, in Of Microbes and Life, edited by Jacques Monod and Ernest Borek. New York: Columbia University Press, pp. 50-5.
Dermitzakis, Emmanouil T. and Andrew G. Clark (2009). “Life after GWA Studies”, Science vol. 326 (Oct. 9), pp. 239-40. doi:10.1126/science.1182009
Dumont, Jacques E., Fréderic Pécasse and Carine Maenhaut (2001). “Crosstalk and Specificity in Signalling: Are We Crosstalking Ourselves into General Confusion?”, Cellular Signalling vol. 13, pp. 457-63.
Dunker, A. Keith, Christopher J. Oldfield, Jingwei Meng et al. (2008). “The Unfoldomics Decade: An Update on Intrinsically Disordered Proteins”, BMC Genomics vol. 9 (Suppl 2):S1. doi:10.1186/1471-2164-9-S2-S1
Fletcher, Daniel A. and R. Dyche Mullins (2010). “Cell Mechanics and the Cytoskeleton”, Nature vol. 463 (Jan. 28), pp. 485-92. doi:10.1038/nature08908
Grant, Barry J., Alemayehu A. Gorfe and J. Andrew McCammon (2010). “Large Conformational Changes in Proteins: Signaling and Other Functions”, Current Opinion in Structural Biology vol. 20, pp. 142-7.
Gsponer, Jörg and M. Madan Babu (2009). “The Rules of Disorder Or Why Disorder Rules”, Progress in Biophysics and Molecular Biology. doi:10.1016/j.pbiomolbio.2009.03.001
Holdrege, Craig (1998). “Seeing the Animal Whole: The Example of the Horse and Lion”, in Goethe’s Way of Science, edited by David Seamon and Arthur Zajonc. Albany: SUNY Press, pp. 213-32.
Holdrege, Craig, editor (2002). The Dynamic Heart and Circulation, translated by Katherine Creeger. Fair Oaks CA: AWSNA.
Hon, Gary C., R. David Hawkins and Bing Ren (2009). “Predictive Chromatin Signatures in the Mammalian Genome”, Human Molecular Genetics vol. 18, no. 2, pp. R195-R201. doi:10.1093/hmg/ddp409
Hyduke, Daniel R. and Bernhard Ø. Palsson (2010). “Towards Genome-Scale Signalling-Network Reconstructions”, Nature Reviews Genetics vol. 11 (April), pp. 297-307. doi:10.1038/nrg2750
Kant, Immanuel (2000). The Critique of Judgment, translated by J. H. Bernard. Amherst NY: Prometheus Books.
J. H. Keyak, J. H. (2009). “Reduction in Proximal Femoral Strength Due to Long-Duration Spaceflight”, Bone vol. 44, no. 3, pp. 449-53.
Lavelle, Christophe (2009). “Forces and Torques in the Nucleus: Chromatin under Mechanical Constraints”, Biochemistry and Cell Biology vol. 87, pp. 307-22. doi:10.1139/O08-123
Levy, Emmanuel D., Christian R. Landry and Stephen W. Michnick (2010). “Signaling through Cooperation”, Science vol. 328 (May 21), pp. 983-4. doi:10.1126/science.1190993
Lewontin, R. C. (1992). “The Dream of the Human Genome”, New York Review (May 28), pp. 31-40.
Lu, Xin (2010). “Tied Up in Loops: Positive and Negative Autoregulation of p53”, Cold Spring Harbor Perspectives in Biology 2010;2:a000984 (Dec. 9). doi:10.1101/cshperspect.a000984
Mansfield, Kyle D. and Jack D. Keene (2009). “The Ribonome: A Dominant Force in Co-ordinating Gene Expression”, Biology of the Cell vol. 101, no. 3, pp. 169-81. doi:10.1042/BC20080055
Mattick, John S. (2007). “A New Paradigm for Developmental Biology”, Journal of Experimental Biology vol. 210, pp. 1526-47. doi:10.1242.jeb.005017
Mattick, John S. (2009). “Has Evolution Learnt How to Learn?”, EMBO Reports vol. 10, no. 7, p. 665. doi:10.1038/embor.2009.135
Mattick, John S., Ryan J. Taft and Geoffrey J. Faulkner (2009). “A Global View of Genomic Information: Moving Beyond the Gene and the Master Regulator”, Trends in Genetics vol. 26, no. 1. doi:10.1016/j.tig.2009.11.002
Mayer, Bruce J., Michael L. Blinov and Leslie M. Loew (2009). “Molecular Machines or Pleiomorphic Ensembles: Signaling Complexes Revisited”, Journal of Biology vol. 8, no. 9, article 81.
McClintock, Barbara (1983). “The Significance of Responses of the Genome to Challenge”, Nobel lecture (Dec. 8).
Moss, Lenny (2003). What Genes Can’t Do. Cambridge MA: MIT Press.
Rothman, Stephen (2002). Lessons from the Living Cell: The Limits of Reductionism. New York: McGraw Hill.
Russell, E. S. (1930). The Interpretation of Development and Heredity: A Study in Biological Method. Freeport NY: Books for Libraries Press.
Stove, David (1995). Darwinian Fairytales: Selfish Genes, Errors of Heredity, and Other Fables of Evolution. New York: Encounter Books.
Uversky, Vladimir N. (2010). “The Mysterious Unfoldome: Structureless, Underappreciated, Yet Vital Part of Any Given Proteome”, Journal of Biomedicine and Biotechnology vol. 2010, article ID 568068. doi:10.1155/2010/568068
Weiss, Paul (1959). “Cellular Dynamics”, Reviews of Modern Physics vol. 31, no. 1 (Jan.), pp. 11-20.
Weiss, Paul (1962). “From Cell to Molecule”, in The Molecular Control of Cellular Activity, edited by John M. Allen, pp. 1-72. The University of Michigan Institute of Science and Technology Series. New York: McGraw-Hill.
Weiss, Paul (1963). “The Cell as Unit”, Journal of Theoretical Biology vol. 5, pp. 389-97.
Weiss, Paul (1967). “One Plus One Does Not Equal Two”, in The Neurosciences: A Study Program, edited by Gardner C. Quarton, Theodore Melnechuk and Francis O. Schmitt, pp. 801-21. New York: Rockefeller University Press. Reprinted in Weiss 1971, pp. 213-61.
Weiss, Paul (1970). “The Living System: Determinism Stratified”, in Beyond Reductionism: New Perspectives in the Life Sciences, edited by Arthur Koestler and John R. Smythies. New York: Macmillan, pp. 361-400. Reprinted in Weiss 1971, pp. 262-311.
Weiss, Paul (1971). Within the Gates of Science and Beyond: Science in Its Cultural Commitments. New York: Hafner.
Weiss, Paul A. (1973). The Science of Life: The Living System — A System for Living. Mount Kisco NY: Futura Publishing.
White, Robert J. and Andrew D. Sharrocks (2010). “Coordinated Control of the Gene Expression Machinery”, Trends in Genetics vol. 26, no. 5, pp. 214-20.
Zaidi, Sayyed K., Daniel W. Young, Martin A. Montecino et al. (2007). “Mitotic Bookmarking of Genes: A Novel Dimension to Epigenetic Control”, Nature Reviews Genetics vol. 11 (Aug.), pp. 583-9. doi:10.1038/nrg2827
Zhou, Yaoqi, Dennis Vitkup and Martin Karplus (1999). “Native Proteins Are Surface-molten Solids: Application of the Lindemann Criterion for the Solid versus Liquid State”, Journal of Molecular Biology vol. 285, pp. 1371-1375.
This document: https://bwo.life/mqual/genome_5.htm
Steve Talbott :: The Unbearable Wholeness of Beings
