Biology Worthy of Life
An experiment in revivifying biology
RNA: Dancing with a Thousand Partners
Tags: RNA; explanation/biological; explanation/part and whole; form/organismal; holism; inwardness; gene regulation; evolution/as mindless process
Contents
Protein-coding RNAs (messenger RNAs) have been discovered in large communities of mutual regulation. The result of their intricate interplay is an adjustment of gene and protein expression so complex, so fluid, and so continually varying in its details as to blur the distinction between regulators and objects of regulation — and pose insuperable challenges for analysis.
From static to dynamic understanding
A key question is this: when vast numbers of molecules are participating in extended, highly articulated, and directional cellular processes such as cell division or RNA splicing, what keeps them “on-script” when there are a thousand other things they might do — and will do in different contexts? This question is underappreciated due to a lack of imagination in picturing what we already know about how every specific molecular process adapts to the fluid, ever-changing conditions within the organism.
Explaining from the whole to the part
If we are faithful to observation, we recognize that in the organism the larger pattern, or context, should, in the proper sense, be seen as the cause of the the part-processes through which the pattern comes to manifestation. This is the stumbling block for conventional scientists of a materialistic bent. Yet it turns out to be a truth that they implicitly accept and work with all the time.
Sometimes the prerequisite for scientific advance is courage to put a name to the clearly indicated borders of an unknown territory — a territory we know is there and to which we can assign a definite place in the larger scientific scheme of things, but whose nature and characteristic mode of functioning we do not yet understand. The forming agency so obviously at work in the organism is one such “nameable unknown”.
![]()
To judge from popular science reporting, and even from many technical journals, you might think that the most dramatic developments in molecular biology today are particular discoveries — a new way to produce stem cells, a method for altering the genome, a genetic “cause” of some trait, an unexpected evolutionary relation between two species, a possible treatment for cancer ...
But if you are looking for more profound revelations, you will not be content with these headline discoveries. You will instead reflect upon the insistent subtext of the research reports. Then you will recognize in one paper after another — not at all concealed, but not explicitly acknowledged either — the central elements of a new biology more worthy of life, a biology requiring a mode of understanding far removed from the dominant conceptual framework of the past several decades.
One finds, for example, that in once-isolated and sharply focused areas of molecular biological investigation, the focus is rapidly becoming less sharp. Boundaries are becoming more permeable, so that it is difficult to separate one topic from another. Every “classical” function of a molecule or structure or pathway is turning out to be just one of many different and often (at first) seemingly unrelated functions. Every niche is interwoven with other niches, and the play of “causes” and “effects” is more like the flow of a stream with its endless, interpenetrating eddies than an interaction of discrete machine parts1.
That’s why terms such as “network”, “systems approach”, “interconnected”, “combinatorial complexity”, and above all “crosstalk” and “context-dependence” now show up with such striking frequency in technical papers. The take-home message is that we’re witnessing a transformation in the way we must think of organisms — a transformation whose importance far outweighs any particular discoveries.
A molecular web of life
This quiet revolution occurring beneath the public radar was most recently emphasized for me by a paper in Nature entitled “The Multilayered Complexity of ceRNA Crosstalk and Competition” (Tay et al. 2014*). The paper concerns so-called “competing endogenous RNAs”, or ceRNAs. You will recall that RNA molecules in general are produced (transcribed) from DNA by an enzyme, whereupon some of them, known as “messenger RNAs”, or “mRNAs”, are translated into protein.Many other RNAs are non-protein-coding, and these have in recent years been found to participate in a wide range of regulatory functions. Perhaps most prominent among noncoding RNAs are the very short microRNAs, which can typically pair up with any of numerous messenger or other RNAs and alter their fate, commonly either by leading to their chemical degradation or by repressing their translation. The microRNAs themselves are generated from longer RNAs by various elaborate processes.
So, then, what are competing endogenous RNAs? They are not really a distinctive class of RNA, but rather collections of RNAs caught up within a certain kind of mutual “conversation”. The ones most commonly dealt with in the literature so far are messenger RNAs — that is, protein-coding RNAs. But long noncoding RNAs (of which there are many) and various other RNAs also play roles as ceRNAs. As for “endogenous”, it refers merely to the fact that the RNAs at issue are native to the cells or organisms under discussion, as opposed to being artificial constructs introduced by engineers.
We come to the crux of the matter with the word “competing”, which is probably not the best choice. It would be better to say “mutually regulating” or “mutually influencing”. In any case, an important point is that mRNAs and long noncoding RNAs are often densely packed with binding sites for microRNAs. Moreover, a single such RNA may contain not only multiple binding sites for a specific type of microRNA, but also for different microRNAs; and at the same time a single type of microRNA may bind to any of multiple different ceRNAs.
This luxuriance of interaction partners opens up possibilities for remarkably complex networks where direct and indirect influences are operating in all directions. The basic idea is this: if multiple ceRNAs can be bound by the same microRNAs, then increased expression of one of those ceRNAs may enable it to act as a “sponge”, soaking up the microRNAs and preventing them from binding to and down-regulating the expression of other ceRNAs. This will typically lead to decreased protein production from the sponge (if it is a protein-coding RNA), and increased protein production from any ceRNAs that have been spared repression by the now bound microRNAs.
Of course, to speak of the increased or decreased production of a protein is a huge simplification. If, for example, the protein happens to be one of more than a thousand human transcription factors, each of which may bind to and variously alter the expression of scores or hundreds of genes, and if many of those genes in turn give rise to proteins or regulatory RNAs with their own intricate and far-reaching role within the organism ... well, then, any talk about the effect of this or that molecular transaction begins to look hopelessly provincial.
Given the overall huge number of microRNA binding sites scattered over the RNAs produced from our genomes, the potential for these RNAs to modulate the roles of each other is all but incomprehensible. A change in the expression level of one mRNA may modulate the level of who knows how many others, which in turn will affect all the bodily functions in which those others are implicated. Many potential pathways of interaction circle around (via few or innumerable steps) so that previously influenced members of the network can in turn influence the influencers.
Interpenetrating spheres of influence. Competing endogenous RNAs have gained attention only during the past three or four years, and efforts to characterize them are in an early stage. The entire topic has to date been a minor sideshow, secondary to the main arena where much more high-profile processes bearing on gene expression are analyzed and dissected. But this only makes more significant the fact that such a great regulatory potential is now being recognized in ceRNAs.
The illustration below characterizes one network of interactions, as it has been explored so far. The ceRNAs are shown in orange boxes, with directly “competing” pairs located at opposite ends of the arrows. The microRNAs mediating the competition are listed alongside the arrows.
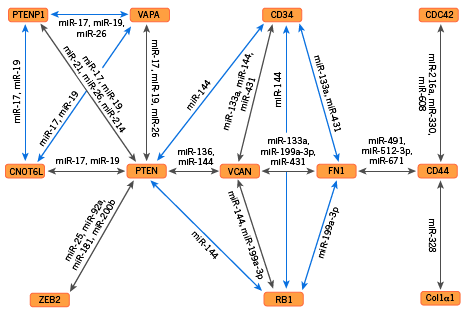
Network of competing endogenous RNAs (orange boxes). Gray arrows show where specific microRNAs are known to bind distinct ceRNAs, and blue arrows indicate possible interactions involving microRNAs known to be present. At least a dozen other ceRNAs not shown here have been identified as involved in the network, but the specific microRNAs mediating their participation have not yet been identified. (From Tay et al. 2014*.)
Let’s consider just two of the RNAs in the figure. PTEN is produced from the PTEN gene and yields a protein (also called “PTEN”) when translated. The protein is, among other things, a tumor-suppressor. In performing this role it conduces to cessation of cell division, often followed by a carefully orchestrated cell death. (In other roles the PTEN protein appears to facilitate cell migration, and to play a part in the adhesion of cells to each other.)
PTENP1, on the other hand, is a so-called “pseudogene”, presumably resulting evolutionarily from a mutational duplication of PTEN, followed by further mutations compromising its protein-coding function. Pseudogenes are one more example of those many DNA elements, once written off as nonfunctional “junk”, which are now being “caught in the act” playing important roles.
In the present case, we know at least one role for PTENP1. Its RNA may be incapable of being translated into protein, but it nevertheless shares many microRNA binding sites with the RNA of its progenitor gene, PTEN. By sequestering those microRNAs away from PTEN, PTENP1 allows the tumor-suppressor to be expressed at proper levels. If, on the other hand, the pseudogene becomes dysregulated for some reason, then microRNAs that would otherwise bind to PTENP1, end up instead binding to, and repressing, PTEN, which reduces its tumor-suppressing activity. It has in fact been shown that PTENP1 functioning is selectively lost in human cancers, consistent with its importance as a microRNA sponge.
Each RNA in the network depicted here has its own story to relate within the organism — and so, too, each microRNA. The diagram shows only relatively few interactions — though more than enough to make for a complexity impossible to track in any precise way. The authors refer to a study of brain cancer (glioblastoma) where “the analysis was significantly extended beyond the binary ceRNA associations described in most other studies”, and “the PTEN ceRNA interactions were found to be part of a post-transcriptional regulatory layer comprising more than 248,000 microRNA-mediated interactions2”.
The end of a simplistic formula. Perhaps the most unexpected aspect of all the above lies in the discovery that protein-coding RNAs (mRNAs) are not only the intense focus of massive and ubiquitous regulatory activity directed at them from all sides, but many mRNAs are at the same time engaged in regulatory activity of their own, correlated with the rise and fall of their individual expression levels. Thousands of mRNA (and other) transcripts can, in the words of Tay et al., “communicate and regulate each other by competing specifically for shared microRNAs”.
It is clear, then (if one didn’t already recognize this!) that mRNAs are not just an intermediate element in a straightforward assembly line for producing protein, as the traditional formula would have it:
Rather, they participate in a vast “breathing” process, one of whose primary outcomes is the regulation — the balancing and counterbalancing — of the mRNAs themselves in proper relation to ever-changing conditions in the cell and its environment. The challenge for our understanding is considerable when we realize that all these RNAs are (to revise the metaphor only slightly) “swimming” in a common pool, one whose significant eddies and currents can be intricately distinct even as they continually flow one into the other.
The upshot of it all is that protein-coding RNAs are found to share broadly in the fluid life of the organism, and not to be mere cogs in a deterministic mechanism. In particular, they gain additional, noncoding (regulatory) functions, and the sharp distinction between coding and noncoding regions of DNA begins to look even more artificial than it has already become.
How, then, should we reconceive the linear formula shown above? The rather too simple answer is that we should picture an (at present) incomprehensible mass of circular, bidirectional, and bifurcating arrows passing in, through, and around each of the three terms, in all directions, and intersecting with everything else going on in the organism. But that doesn’t help much, except so far as we see the essential implications of it. And the implications, in today’s scientific environment, are not easy to get at. But here is my best attempt.
From static to dynamic understanding
I have, in passing, more than once asked this question: When molecules in the thick soup of the cellular plasm cooperate to carry out an extended, highly articulated, directional process such as cell division or RNA splicing, what constrains these molecules to interact in such a sensitive, highly choreographed way, rather than to wander off-script and enter into any of a thousand other perfectly natural molecular engagements?I find it a stunning fact about contemporary biology that this sort of question rarely gets asked. Stunning because we have no good answer for it, and without an answer we have little understanding of the life of the organism. Unfortunately, however, the question easily disappears from view owing to a failure of imagination — that is, a failure to picture as broadly and concretely as possible, and in full detail, how relationships within the organism play out over time and under varying conditions.
Consider the drawing above. Its elements do not represent a set of fixed entities in fixed relationships. The whole point, rather, has to do with changing relationships. But even before looking at the dynamism delimited by the box of the illustration, we need to recognize that each one of the depicted mRNA and microRNA molecules has its own unthinkably complex “life narrative”3 outside the box, through which it has come to be where it is now. It has its own regulated patterns of initial synthesis, typically with variations on different occasions, followed in any case by function-altering chemical modifications, and with complexly regulated conditions for degradation — these latter also being decisive for its proper functioning. So the kaleidoscopically shifting relationships suggested in the drawing are actually relationships between narratives, not between static things.
But after making some effort to imagine all that, we still have only a relatively static image of the living processes at work. The entire network of relationships shown in the drawing — the collective performance of the entire group — must mesh with innumerable other molecular pathways and activities comprising the organism’s life and often involving some of the same molecules. None of these pathways and activities are mere givens, but are changing and never fully predictable expressions of the organism’s unique life trajectory and encounter with its environment.
Clearly, then, the relationships made visible in drawings such as the one above, complex as they may seem, nevertheless reduce reality almost to a nullity. In order to flesh out that reality just a little, here are a few relevant considerations:
![]() What stage of the cell cycle are the molecules participating in? Just
about everything becomes different when the cell passes from a resting
stage to the stage where DNA is replicated, and again when the cell goes
through
mitosis
(cell division). The activity of billions of molecules is redirected in a
coordinated way. In other words, some of the potentially errant
performances that once would have been (as I put it above) “off-script”
now appear to be just what the script calls for. This raises the obvious
question: how are the innumerable molecules assembled and organized
according to the right overall script — or, rather, the many simultaneous
scripts and sub-scripts constituting the integral life of the organism?
What stage of the cell cycle are the molecules participating in? Just
about everything becomes different when the cell passes from a resting
stage to the stage where DNA is replicated, and again when the cell goes
through
mitosis
(cell division). The activity of billions of molecules is redirected in a
coordinated way. In other words, some of the potentially errant
performances that once would have been (as I put it above) “off-script”
now appear to be just what the script calls for. This raises the obvious
question: how are the innumerable molecules assembled and organized
according to the right overall script — or, rather, the many simultaneous
scripts and sub-scripts constituting the integral life of the organism?
![]() What kind of cell are we talking about? The molecules shown in the
drawing will integrate with larger patterns of activity that drastically
differ between a red blood cell and a brain neuron or muscle cell.
What kind of cell are we talking about? The molecules shown in the
drawing will integrate with larger patterns of activity that drastically
differ between a red blood cell and a brain neuron or muscle cell.
![]() What stage of
development
is the organism going through, and what stage of
differentiation
characterizes the particular cell of interest? Also, what is the age of
the cell? The age of the organism? Patterns change with development and
age.
What stage of
development
is the organism going through, and what stage of
differentiation
characterizes the particular cell of interest? Also, what is the age of
the cell? The age of the organism? Patterns change with development and
age.
![]() What illness or pathological condition might this particular cell be
affected by? Is it showing an inflammation response? Or is it subject to
some particular stress, such as a mechanical stress or physical injury or
heat shock? Has it been subjected to damaging radiation, so that a
DNA
damage response — or perhaps an orderly cell death — needs to be arranged?
In all such cases, the cell “marches to a different tune” as required by
circumstances, which means that untold numbers of molecular trajectories
need to be integrated in new and dynamically varying ways.
What illness or pathological condition might this particular cell be
affected by? Is it showing an inflammation response? Or is it subject to
some particular stress, such as a mechanical stress or physical injury or
heat shock? Has it been subjected to damaging radiation, so that a
DNA
damage response — or perhaps an orderly cell death — needs to be arranged?
In all such cases, the cell “marches to a different tune” as required by
circumstances, which means that untold numbers of molecular trajectories
need to be integrated in new and dynamically varying ways.
![]() What is the organism as a whole doing? Growing, resting, running,
sleeping, smelling danger, hibernating, playing, courting? Everything in
the organism “tries” to be consistent with its current activity.
What is the organism as a whole doing? Growing, resting, running,
sleeping, smelling danger, hibernating, playing, courting? Everything in
the organism “tries” to be consistent with its current activity.
There is, of course, much more if we would come fully to terms with the real-life complexity, diversity of performance, varied narrative threads, and integration of narratives that we see everywhere in the organism. Moreover, we see these things without the sort of consistent causal linkage from one state to another typical of machinery or of computers executing software routines. All those molecules really could wander off according to different scripts, and in fact are always doing so — but in a well-gauged relation to changing circumstances. That is, they do so in a way that meaningfully holds together. Their consistency is not mechanical, but rather is the consistency required for a dynamic and unique narrative — the narrative of one organism’s life as an exemplar of its species.
Evolution and belief
Biologists frequently express their amazement at the number of Americans who, according to the polls, do not “believe in evolution”. There are, no doubt, all sorts of poor reasons for such a stance. But few scientists seem to recognize how little choice the polled individuals are given. As biologists have presented the matter to them, a “yes, I believe” answer commits them to an unbearable lie — namely the lie that everything I have been describing above is not what it so obviously shows itself to be. It is not, they are told, a story of currently unfathomable wisdom and profoundly meaningful and well-directed (purposive) activity, but rather a story of blind, undirected, meaningless forces. To assent to such a belief would be to deny the evidence of their own eyes — a denial biologists may have become comfortable with thanks to the consolation of their philosophical prejudices4.
But we can be thankful that so many people, both educated and not so educated, refuse to share those prejudices; and even when deprived of the scientific community’s help in making proper sense of highly technical issues, they still have a sound enough intuition to rebel against patent nonsense. If one consequence of this rebellion is the rejection of evolution along with the lie — well, we can hope that the resulting cultural impasse will eventually drive some scientists to look again at the most fundamental questions raised by their work. Then they may be able, with a little humility, to help educate the general public in a liberating, rather than an arrogant and oppressive, manner.
And this is where the failure of imagination occurs. The explanations we are typically handed from the laboratories of molecular biology never touch the holding together of the narrative, apparently because biologists are not keeping the problem in view as they isolate a particular interaction and say, “Ah, now we know that this makes that happen in such-and-such a way”. They are absolutely right, of course, but are forgetting that what happens in such-and-such a way will very likely happen in a different way under changing circumstances. And that is what biologists need to address if they are to become as coherent in their understanding as the organism is in its being. (See also accompanying box.)
Because the philosophical bias toward an inorganic style of explanation is so strong in biology, as soon as a single step in an organic process has been “mechanistically” elucidated (as the usual phrasing goes), the biologist’s thinking tends to come to a stop, satisfied that the only thing required is to add more such explanations, step by step. But no such piling up of explanations gets us to the organicity of the process, with all its power of twisting and turning in a coherent way. What is captured, rather, is merely a snapshot of what we might approximately think of as the causal relations at one moment, in one set of circumstances. But the power of the organism’s life is a power to reorder all those causal relations as part of an ongoing narrative.
Explaining from the whole to the part
The usual modes of biological explanation are ways of showing, or attempting to show, how one thing precisely determines another, whether logically, algorithmically, or mathematically (Talbott 2004*). This quest for an exact, replicable determination of results, often associated in one way or another with the notion of “causal mechanisms”, is the product of one-sided, analytical thinking that can never lay hold of life.We can rightly take pride in our powers of analysis. The picture of ceRNAs, of which I tried to give some inkling above, would never have come within our ken if there had been no stringent analyses isolating the elements of complex processes. But we cannot simply recombine those elements and, by calling them “causal factors”, reach the narrative reality that initially struck us as something worth trying to understand.
Truly biological problems, as opposed to physical or chemical ones, are always first posed for us at a narrative level, and translating them into a collection of local problems of cause and effect never gets us back to the problems we began with. Gaining some appreciation for this fact is crucial to the future of biology.
A good way to start is to recall how twentieth-century cell biologist Paul Weiss referred to the relation between part and whole. (See the discussion of Weiss’ work in my article section, “What Does ‘More Than the Sum of Its Parts’ Mean?” — Talbott 2010*.) Weiss demonstrated with numerous examples that the many degrees of freedom exhibited by low-level processes are increasingly constrained at a higher level, so that the larger-scale results are more “deterministic” than the fine-scale details can possibly explain:
A cell retains its identity far more conservatively and remains far more similar to itself from moment to moment, as well as to any other cell of the same strain, than one could ever predict from knowing only about its inventory of molecules, macromolecules, and organelles, which is subject to incessant change, reshuffling, and milling of its population (Weiss 1973, p. 25*).
For more on this, see my earlier discussion. The clear facts of the matter directly confute any conviction that we can explain the organism from the bottom up5.
Weiss is not, however, suggesting that there is machine-like determinacy at the level of the whole, or that the organism is any less the adaptable and flexible being we know it to be. It’s just that (to offer my own take on his point) a disciplining of lower-level activity occurs from above — not in a conventional (efficient) causal sense, but in the sense that a larger-scale pattern, an intentionality, shapes the various part-processes through which that very pattern and intention become manifest.
This shaping discipline occurs despite all kinds of apparently incidental variations in the details — despite, that is, a degree of low-level freedom inconsistent both with the idea of strict rule-following and with the conviction that we could predict the whole from the lawfulness or behavior of the parts, seen merely as parts. The parts become revelatory only insofar as we see them participating in, and deriving their life from, the whole. And that whole is not some static form, but rather the dynamic story, laced with contingencies, of a recognizable organism.
This kind of thing is easy to picture in terms of the ceRNAs discussed above. You would doubtless have a very difficult time predicting the interactions of individual molecules; they are not in fact conforming to any set of fixed rules. But in the case of all such complex interactions in the organism, our growing knowledge of the larger story enables us to make sense of the “general drift” of the collective molecular processes. And this general drift is neither more nor less than the meaning imparted to the details by the storytelling agency: the organism.
As I have emphasized in many articles, every organism’s life is a narrative — a fact reflected in the context-dependence of every detail. The context is the larger narrative that gives proper force and direction and meaning to the particular “words”. But this is to say that, whatever else it may be or do, the organism presents itself as a kind of narrative text — it manifests itself as an ongoing embodiment of thought. Certainly, for the bacterium or gazelle, this is not thought it has become aware of, or “made its own” in anything like the human sense. But we don’t need evidence of self-awareness in order to recognize the play of thought, whether we’re watching an insect weave a cocoon prior to metamorphosis, or are reflecting on the mathematical elegance of the heavens.
Courage to name the unknown
The organism as “a being of thought” — there’s the rub for scientists in our day. When I remark, as above, that in the organism larger-scale patterns shape the various part-processes through which those very patterns and intentions become manifest, I know the incredulous response very well. How can something that is not “there” — something we cannot “finger and peep at” (in Coleridge’s phrase) — bring things to appearance?And yet, the incredulity is ill-considered, because I am only emphasizing what every scientist implicitly believes: an important form of causation runs from the whole — a whole that must at least in part be understood as a body of thought — to the parts through which the whole manifests itself.
This is true even at the inorganic level. When confronted with two gravitational masses, we may be inclined to think of one mass attracting the other. One is the actor, the other is acted upon. But the physicist knows very well that this isn’t right. The two bodies are never observed in anything but a mutual, inseparable, and irreducible embrace. Yes, we can say that A attracts B with a particular force, and at the same time B attracts A with a particular force, and we can draw two distinct vectors on our force diagram. But this analysis is for purposes of calculation only, and doesn’t tell us that there are two separable forces involved — doesn’t in fact give us much understanding at all of what is involved, which remains pretty much the mystery for us today that it was for Newton in his day, even if our analytical and calculational powers have increased astronomically.
So from the solar system to distant galaxies we see, among other things, gravitation-related patterns , and we can capture certain lawful aspects of these diverse patterns with mathematical formulae. The patterns and formulae, as elements contributing to our understanding, are nothing if not conceptual, or ideational. Laws and relations are ideas, not things. Physicists cannot help believing that everything new coming to manifestation in the solar system will illustrate the consistent thought-relations they know before the fact as celestial mechanics6.
This kind of patterning may be very different from what we observe in organisms — it is not, for example, in the same way a storytelling — but it nevertheless illustrates how something we necessarily characterize as a thought-structure must be seen as somehow shaping the very phenemona through which we come to recognize the thought-structure.
In the case of the organism, the patterning, or forming, activity shows itself at work in every experimental context — including that of competing endogenous RNAs. And we can understand that the various entities caught up in that activity do not themselves explain it — cannot even become themselves apart from it. I will not here propose a special name for this forming activity; there are plenty of (more or less problematic) names to choose from historically, if we should so wish. In any case, the important thing is to acknowledge the activity, whatever we may call it, as a severe lacuna in our biological understanding.
The good news is that the lacuna, once we acknowledge it, does not prevent our continuing to advance in understanding — and in particular to recognize with ever greater clarity the ideal, qualitative, forming activity taking place before our eyes, and to characterize it in all the detail we can7. In the end, I suspect, it is this faithful characterization that will bring us toward the fuller understanding we seek.
Notes
1. There are, to be sure, hierarchical and modular relationships, especially in more complex organisms. These are critical for our understanding. But the “modules” are not machine modules. They could better be compared to the more or less distinct vortices within a stream, which are context-dependent, lack precise boundaries, and gracefully transition one into the other.
2. There are complexities in every direction. To pick a few at random:
- A significant portion of human microRNAs undergo RNA editing before becoming fully functional. This can drastically change their performance, or even lead to their degradation. Nothing of that realm of regulation is indicated in the diagram.
- Looking at just one of the other RNAs in the diagram: “Interestingly, while the ZEB2 ceRNA opposes melanoma by sponging the microRNAs that would otherwise repress PTEN’s tumor suppression activity, the ZEB2 protein is known to promote other cancers. ‘It is astonishing that RNA and protein molecules encoded by the same gene can take part in opposing biological processes’, [postdoctoral fellow Florian] Karreth said” (BIDMC 2011*).
- And on pseudogenes: “The relationship between pseudogenes and their parental counterparts [that is, the genes from which they originated by duplication] is extremely varied. The pseudogene can promote or inhibit the expression of or enhance or impair the function of the parental gene. It is conceivable that the function of the parental gene is impaired by the pseudogene at one level (for instance, the pseudogenic protein is an allosteric inhibitor of the protein encoded by the parental gene) but is promoted at another level (for instance, the pseudogenic RNA competes with the parental RNA for a microRNA). Pseudogenes could thereby act as sophisticated regulators of their parental counterparts, finely tuning every step of the parental genes’ expression as well as their activity” (Poliseno 2012*).
- Or, if you dare to browse some notes suggesting the real extent of the interconnections as an organism manages its DNA, see How the Organism Decides What to Make of Its Genes.
3. It would be a useful exercise to browse through some notes relating to at least one facet of this narrative — for example, the alternative splicing of mRNAs through which any given protein-coding gene can yield many different RNAs coding for up to hundreds of distinct proteins.
4. Unfortunately, this kind of denial is often explicit or implicit in arguments we see coming from the resurgent community of intelligent design theorists (as I pointed out in “A Sectarian Quarrel? Intelligent Design and Neo-Darwinism”). Rather than seeing an intelligence at work in the organism as a living capacity, they would rather reserve such intelligence for God, with the organism reduced to a kind of machine that God may occasionally tinker with (miraculously, it would seem) from without. Nevertheless, their critiques of conventional evolutionary theory have grown increasingly sharp and on-target in recent years. I hope to have more to say about all this shortly.
5. Weiss referred to “microindeterminacy” and “macrodeterminacy”. Here is some of what he had to say about these terms: (Weiss 1973*):
“A macroevent can be fully determinate in the sense that, given the premises, we can predict the general outcome, the confidence of our prediction resting on the infallibility of countless earlier experiences; yet at the same time, the component microevents involved might take courses that are in detail absolutely unpredictable and unique — unpredictable, because being unique and nonrecurrent, they have had no exact precedent in our experience. ... To me, the main flaw in judging ‘determinism’ lies in its equating science with the doctrine of microprecise causality, or ... microdeterminism” (pp. 17-8).
“While definite regions of the early embryo are predictably the rudiments for the formation of heart, liver, kidney, brain, etc., the individual fates of the cells derived from those areas are still unsettled. In other words, overall regularity in the gross is attained and maintained not as a mechanical result and a reflection of a corresponding underlying regularity of rigidly stereotyped behavior of the component elements down to the smallest detail, but on the contrary, in spite of a high degree of vagrancy among the latter. The individual component does not ‘know’ where and in what specialty it will end up until it has been well on its way and given a defining ‘cue’. The cue will vary in accordance with the position the part assumes in the domain of the respective subsystem; and in dynamic terms, geometric location simply denotes the constellation of forces and processes at that point” (pp. 24-5).
6. I am not at all suggesting that current conceptions of celestial mechanics are complete or adequate for understanding the celestial phenomena they are intended to address. For a nonstandard view see holoscience.com.
7. For the kind of qualitative and contextual exploration that can meaningfully situate more narrowly focused analytical investigations, see, for example, the whole-organism studies by my Nature Institute colleague, Craig Holdrege: on the sloth (1999*), elephant (2003*), giraffe (2003*), and skunk cabbage (2000*). The full text of each of those studies is freely available via the indicated links.
Sources:
BIDMC [Beth Israel Deaconess Medical Center] (2011). “Scientisits [sic] Uncover Evidence of Powerful New Cancer Genetic Networks”, press release (Oct. 14). Available online here.
Holdrege, Craig (1999). “What Does It Mean To Be a Sloth?” NetFuture #97 (Nov. 3). Available at https://natureinstitute.org/nature/sloth.pdf.
Holdrege, Craig (2000). “Skunk Cabbage”, In Context #4 (fall), pp. 12-18. Available at https://natureinstitute.org/pub/ic/ic4/skunkcabbage.pdf.
Holdrege, Craig (2003). “The Flexible Giant: Seeing the Elephant Whole”, Nature Institute Perspectives no. 2. Available at https://natureinstitute.org/pub/persp/1/details/eleph.htm.
Holdrege, Craig (2005). “The Giraffe’s Long Neck: From Evolutionary Fable to Whole Organism”, Nature Institute Perspectives no. 4. Available at https://natureinstitute.org/pub/persp/4/.
Poliseno, Laura (2012). “Pseudogenes: Newly Discovered Players in Human Cancer”, Science Signaling vol. 5, no. 242, re5 (Sep. 18). doi:10.1126/scisignal.2002858
Talbott, Stephen L. (2004). “The Reduction Complex”, NetFuture #158 (Nov. 9). Available at http://BiologyWorthyofLife.org/mqual/ch04.htm
Talbott, Stephen L. (2010). “The Unbearable Wholeness of Beings”, The New Atlantis no. 29 (fall), pp. 27-51. Original version published in NetFuture #181 (Dec. 9, 2010). Available at http://BiologyWorthyofLife.org/mqual/genome_5.htm
Tay, Yvonne, John Rinn and Pier Paolo Pandolfi (2014a). “The Multilayered Complexity of ceRNA Crosstalk and Competition”, Nature vol. 505 (Jan. 16), pp. 344-52. doi:10.1038/nature12986
Weiss, Paul (1973). The Science of Life: The Living System — A System for Living. Mount Kisco NY: Futura Publishing.
Further information: See the section “From differences to character” in my article, “Missing Heritability — Or Whole-Organism Inheritance?”: http://BiologyWorthyofLife.org/mqual/genome_10_inherit.htm#gene_diffs
This document: https://bwo.life/org/comm/ar/2014/rna-partners_16.htm
Steve Talbott :: RNA: Dancing with a Thousand Partners

